Research Directions:
Environmental genomics
Marine biodiversity
Bioinformatics and biodiversity informatics
Description:
 Microbial community profiling of marine environments through metagenomics has the potential to transform our understanding of the ecosystem services supported by microbial organisms. Moreover, microbiome diversity is directly correlated with environmental quality through the food chain and the physical and chemical effects of secondary metabolites. Many of the latter provide targets for bioprospecting in medicine and industry.
Microbial community profiling of marine environments through metagenomics has the potential to transform our understanding of the ecosystem services supported by microbial organisms. Moreover, microbiome diversity is directly correlated with environmental quality through the food chain and the physical and chemical effects of secondary metabolites. Many of the latter provide targets for bioprospecting in medicine and industry.
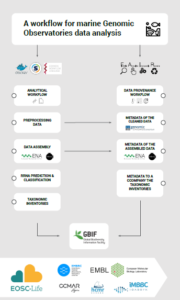
The project will design an effective workflow and deployment strategy for the analysis of the metagenomic genomic observatory (GO) data. It is considered of great importance that the data and preliminary results are made available rapidly to the marine biological community, but also that the data are quality controlled and standardised before release.
The proposed work is aimed at making the large volumes of data produced by the GOs more easily interpretable by the marine biological community by providing the taxonomic inventories of each sample in a timely manner and in a non-technical format. This way, researchers without the technical expertise to analyse such data on their own, as well as policy makers and management agencies, will have the opportunity to inspect the results and understand the biodiversity patterns that are being shaped by the marine microbial communities.






