Environmental Microbiology Lab
Key research areas
Metagenomics, Environmental genomics, Marine biodiversity, biotechnology, Microbial phylogenetics and phylogenomics, Microbial evolutionary genomics, metabarcoding, Environmental status of marine ecosystems, ecology and ecosystem management, Ecological data analysis
People involved

All regions of Earth’s biosphere including the waters of Earth’s oceans, the soil beneath our feet and even the air we breathe teem with microbes. Microbial communities play a significant ecological and biogeochemical role in marine ecosystems, and represent a valuable reservoir of genetic resources with potential applications in biotechnology, ecosystem management and bioremediation. With the rise of molecular genetic tools, it has become apparent that only a very small fraction of the microbial world has been documented, and an even smaller one characterized. Despite the vital role of microorganisms in marine ecosystems and their potential biotechnological applications, little is known about their diversity, ecology and metabolic capabilities.
The Environmental Microbiology Laboratory of IMBBC is designed to investigate microorganisms including Bacteria, Archaea and Fungi from a wide range of marine environments in the context of their diversity, ecology and biotechnology potential. We mostly focus on the exploration of the unique Mediterranean Sea, which is one of the most diverse environments on Earth.
We follow a multidisciplinary approach, using state-of-the-art technologies, molecular-based methods, traditional culture-based methods and computational biology to study the full spectrum of microbial diversity and corresponding key processes.
Our activities:
1) Life in Extreme Environments: Exploring Volcanologically Active Marine Environments and Submarine Caves and Lakes
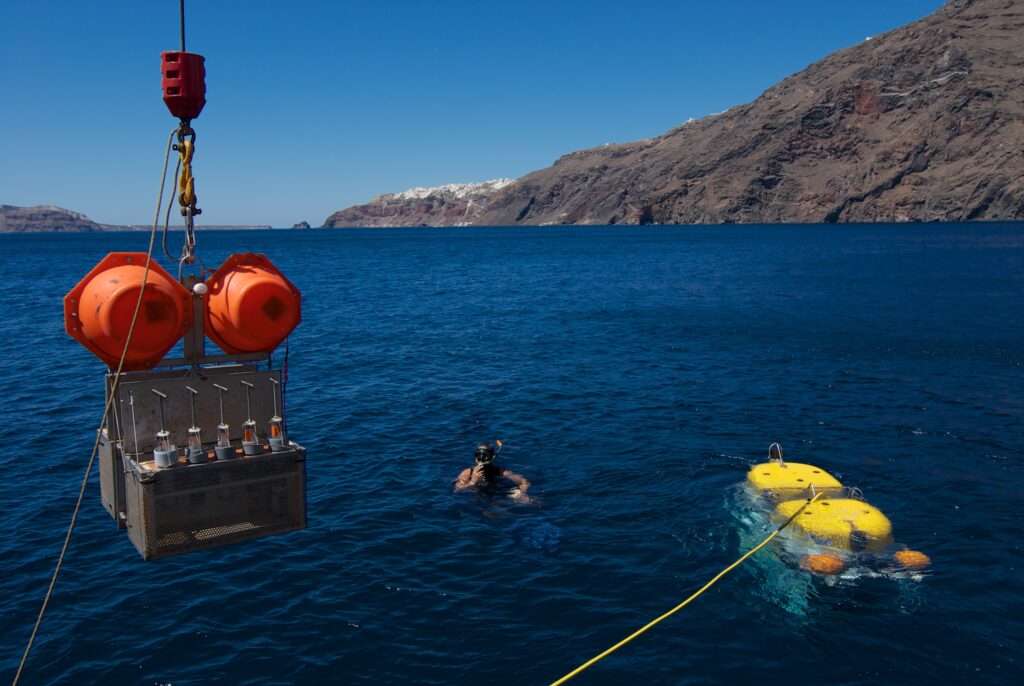
 Microbes that make up most of the Earth’s biomass, can survive in some of the most extreme conditions on the planet; they thrive in extreme hot niches at 122 °C, in frozen sea water at −20 °C, in salt solutions, and in acidic (pH=0) and alkaline environments (pH=12.8). Microorganisms have been even discovered several kilometers below the surface of the Earth’s crust and at high pressures of up to 110 MPa comprising the extreme “sub-seafloor biosphere”. We conduct microbiological research on extreme environments, mainly on the extreme environments of the Hellenic Volcanic Arc. Our work is based on the organization and implementation of sampling expeditions using oceanographic research vessels in extreme environments including submarine volcanoes, hydrothermal vents, active and inactive polymetallic chimneys, submarine caves and lakes etc.
Microbes that make up most of the Earth’s biomass, can survive in some of the most extreme conditions on the planet; they thrive in extreme hot niches at 122 °C, in frozen sea water at −20 °C, in salt solutions, and in acidic (pH=0) and alkaline environments (pH=12.8). Microorganisms have been even discovered several kilometers below the surface of the Earth’s crust and at high pressures of up to 110 MPa comprising the extreme “sub-seafloor biosphere”. We conduct microbiological research on extreme environments, mainly on the extreme environments of the Hellenic Volcanic Arc. Our work is based on the organization and implementation of sampling expeditions using oceanographic research vessels in extreme environments including submarine volcanoes, hydrothermal vents, active and inactive polymetallic chimneys, submarine caves and lakes etc.
2) Monitoring and Sampling Extreme Environments: Demonstration of intelligent, multi-purpose autonomous marine vehicles
Within the framework of the new MERLIN programme (Horizon Europe), our laboratory actively contributes to the development and deployment of next-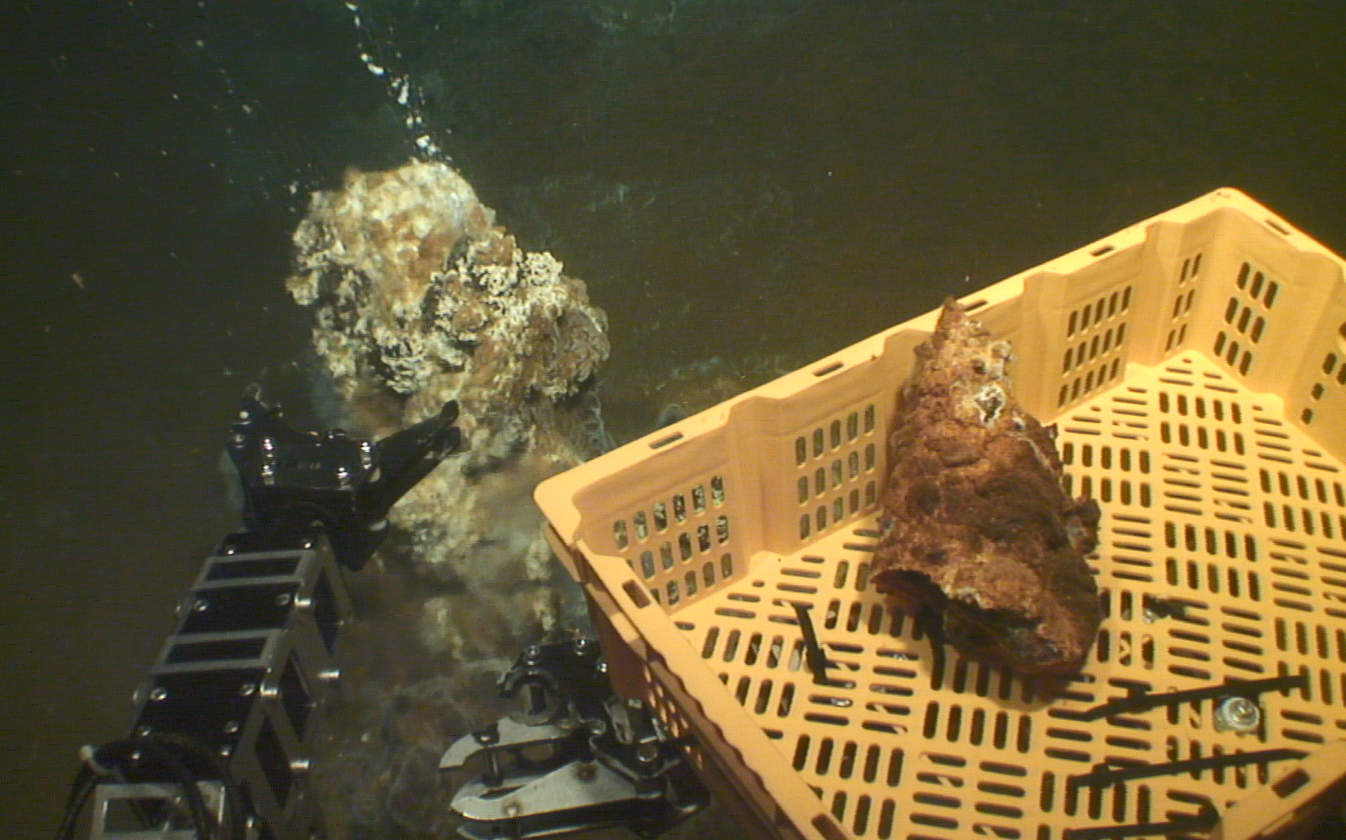 generation autonomous robotic systems for marine exploration. In this context, we are advancing automated sampling and monitoring of extreme submarine volcanoes, by integrating intelligent unmanned surface (USV) and autonomous underwater vessels (AUVs) capable of long-endurance autonomous operation. The new technology will be demonstrated during a major oceanographic expedition, that will be organized and conducted by the IMBBC at the Kolumbo submarine volcano, the most active submarine volcano in the Eastern Mediterranean Sea.
generation autonomous robotic systems for marine exploration. In this context, we are advancing automated sampling and monitoring of extreme submarine volcanoes, by integrating intelligent unmanned surface (USV) and autonomous underwater vessels (AUVs) capable of long-endurance autonomous operation. The new technology will be demonstrated during a major oceanographic expedition, that will be organized and conducted by the IMBBC at the Kolumbo submarine volcano, the most active submarine volcano in the Eastern Mediterranean Sea.
3) Hypersaline Microbial Mats
 Microbial mats are vertically stratified communities of functional groups of microorganisms embedded in an organic matrix, which may also contain minerals such as silicat es and carbonates. They grow on a solid substrate (e.g. sand) and the vast majority of microbial mats is utilizing inorganic carbon as carbon source, hence it is autotrophic. Microbial mats are characterized by pronounced physiochemical gradients which allow for the presence of high species diversity, encompassing a wide range of metabolic capabilities; thus, mats are ideal models to study a whole ecosystem and are considered as natural laboratories. These physicochemical gradients provide microenvironments for various microbial functional groups, which exhibit a certain physiology with which they fulfill a specific function. Although microbial mats have received attention from the scientific community, the full spectrum of their microbial processes and metabolic capabilities has yet to be revealed. We are investigating the microbial diversity and function of hypersaline swamp sediment, where the formation of microbial mats are observed.
Microbial mats are vertically stratified communities of functional groups of microorganisms embedded in an organic matrix, which may also contain minerals such as silicat es and carbonates. They grow on a solid substrate (e.g. sand) and the vast majority of microbial mats is utilizing inorganic carbon as carbon source, hence it is autotrophic. Microbial mats are characterized by pronounced physiochemical gradients which allow for the presence of high species diversity, encompassing a wide range of metabolic capabilities; thus, mats are ideal models to study a whole ecosystem and are considered as natural laboratories. These physicochemical gradients provide microenvironments for various microbial functional groups, which exhibit a certain physiology with which they fulfill a specific function. Although microbial mats have received attention from the scientific community, the full spectrum of their microbial processes and metabolic capabilities has yet to be revealed. We are investigating the microbial diversity and function of hypersaline swamp sediment, where the formation of microbial mats are observed.
4) Μarine Fungi
 Μarine fungi are an untapped source of chemical and taxonomic diversity. This is due to their high potential in decomposing different sources of material, both organic and synthetic, and the limited sampling of marine representatives from this kingdom. It is only in recent years that we have come to appreciate their roles in marine environments, often mediated through symbioses with important ecosystem players, such as sponges and corals. Combining access to emerging -global and local- fungal collections, and high-performance computing facilities, we cultivate and generate annotated genomes and transcriptomes of fungal isolates from diverse habitats. Τhese data are used to investigate symbiosis and other fungal life history traits such as reproductive mode from an evolutionary and functional point of view, adopting molecular evolution and expression profiling approaches. We are interested in the genomic causes and consequences of adaptation to marine environments and search for fungal genes and non-coding elements that are associated with these. Current areas of research include characterization of enzymes for decomposing organic pollutants relevant to applications in bioremediation (collaboration with Biotechnology lab at NTUA-Athens), and taxonomic placement of new isolates (as part of the CMBR project).
Μarine fungi are an untapped source of chemical and taxonomic diversity. This is due to their high potential in decomposing different sources of material, both organic and synthetic, and the limited sampling of marine representatives from this kingdom. It is only in recent years that we have come to appreciate their roles in marine environments, often mediated through symbioses with important ecosystem players, such as sponges and corals. Combining access to emerging -global and local- fungal collections, and high-performance computing facilities, we cultivate and generate annotated genomes and transcriptomes of fungal isolates from diverse habitats. Τhese data are used to investigate symbiosis and other fungal life history traits such as reproductive mode from an evolutionary and functional point of view, adopting molecular evolution and expression profiling approaches. We are interested in the genomic causes and consequences of adaptation to marine environments and search for fungal genes and non-coding elements that are associated with these. Current areas of research include characterization of enzymes for decomposing organic pollutants relevant to applications in bioremediation (collaboration with Biotechnology lab at NTUA-Athens), and taxonomic placement of new isolates (as part of the CMBR project).
5) Coastal Lagoons
 Lagoons are naturally enriched habitats, with unstable environmental conditions caused by their confinement from the sea and their shallow depth. They are particularly vulnerable to human activities and especially to pollution. Such ecosystems are characterized by increased hypoxia and high concentrations of hydrogen sulfide. We are investigating the microbial communities in lagoonal sediments of the Amvrakikos Gulf (Ionian Sea, Western Greece).
Lagoons are naturally enriched habitats, with unstable environmental conditions caused by their confinement from the sea and their shallow depth. They are particularly vulnerable to human activities and especially to pollution. Such ecosystems are characterized by increased hypoxia and high concentrations of hydrogen sulfide. We are investigating the microbial communities in lagoonal sediments of the Amvrakikos Gulf (Ionian Sea, Western Greece).
6) Water Microbiology
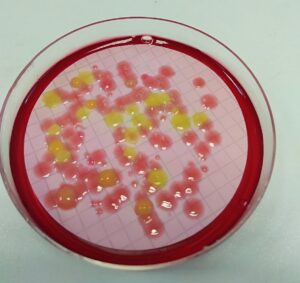 The Water Microbiology Section covers the use of cultivation-based approaches for the detection, identification and enumeration of microorganisms in natural environments with emphasis on the coastal zone (e.g. ports, swimming areas etc). Our lab is fully equipped to undertake the microbiological analysis of water and other environmental samples with high standards. We carry out specific analyses on anthropogenic disease-causing microbes and in particular, coliform, faecal coliform, and streptococci strains. We can further identify unknown microorganisms and classify them to species levels by using molecular-based approaches.
The Water Microbiology Section covers the use of cultivation-based approaches for the detection, identification and enumeration of microorganisms in natural environments with emphasis on the coastal zone (e.g. ports, swimming areas etc). Our lab is fully equipped to undertake the microbiological analysis of water and other environmental samples with high standards. We carry out specific analyses on anthropogenic disease-causing microbes and in particular, coliform, faecal coliform, and streptococci strains. We can further identify unknown microorganisms and classify them to species levels by using molecular-based approaches.
7) The IMBBC Microbial Strain Collection
 Through European and National Research Projects, we have initiated the HCMR-IMBBC microbial strain collection from extreme and highly oligotrophic environments. Our strain collection is continuously being enriched with isolates from various marine environments from the E. Mediterranean Sea including the extreme environments of the Hellenic Volcanic Arc for further exploitation into different biotechnological sections.
Through European and National Research Projects, we have initiated the HCMR-IMBBC microbial strain collection from extreme and highly oligotrophic environments. Our strain collection is continuously being enriched with isolates from various marine environments from the E. Mediterranean Sea including the extreme environments of the Hellenic Volcanic Arc for further exploitation into different biotechnological sections.
In addition, the Microbiology Lab is used for studying Fungi either as marine taxa symbionts (e.g. of sponges) or as free-living organisms. Expertise in culturing and genetically manipulating Fungi is combined with access to global sampling isolates through collaborations and computational mining of multi-omic data.
Related Content
-
Three new papers from the largest oceanographic expedition ever conducted in the Eastern Mediterranean, the Hellenic Arc Volcanic Field IODP Expedition 398, with the participation of IMBBC, have been published in the leading journals Science Advances, Earth and Planetary Science Letters, and Geophysical Research Letters. These studies shed new light on the geological, volcanic, and climatic past of the South Aegean, revealing how large explosive eruptions and tectonic extension have shaped the region over hundreds of thousands of years.
Full stories in the following links:
https://www.science.org/doi/10.1126/sciadv.ads9642
https://www.sciencedirect.com/science/article/pii/S0012821X25004315
https://agupubs.onlinelibrary.wiley.com/doi/10.1029/2025GL115919
- Our multiomic study of the fungus Aspergillus parasiticus and its enzymes induced during growth in presence of plastic has been published in the journal of Environmental Pollution. As part of the HFRI PlastOmics project, a collaboration between NTUA chemical engineers and HCMR researchers who specialize in fungal omics and bioinformatics, we studied the ability of the fungus to metabolize long-chain alkanes and secrete enzymes that are potentially capable of modifying polyethylene. Read more in the open access article:
https://www.sciencedirect.com/science/article/pii/S0269749125007596
-
Μonitoring the most active submarine volcano: Kolumbo
-
Our last paper in Frontiers in Microbiology about microbial communities in sub-seafloor hydrothermal sediments of the active Santorini-Kolumbo volcanic field
href=”https://www.frontiersin.org/articles/10.3389/fmicb.2023.1188544/full
Polymenakou, Paraskevi N; Nomikou, Paraskevi; Hannington, Mark; Petersen, Sven; Kilias, Stephanos P; Anastasiou, Thekla I; Papadimitriou, Vasiliki; Zaka, Eleutheria; Kristoffersen, Jon Bent; Lampridou, Danai; Wind, Sandra; Heinath, Verena; Lange, Sabine; Magoulas, Antonios. 2023. Taxonomic diversity of microbial communities in sub-seafloor hydrothermal sediments of the active Santorini-Kolumbo volcanic field.Frontiers in Microbiology, 14 pp. 1188544, ISSN: 1664-302X. -
OUR NEW H.F.R.I. PROJECT (COORDINATED BY NKUA)
SANTORini’s seafloor volcanic observatory
-
A new HFRI project coordinated by NΤUA) https://www.chemeng.ntua.gr/plastomics/index.html
-
Our lab will be involved in a n exciting new project investigating the potential of fungi to remove/degrade plastics! The project started this year (2022-2025) and involves A. Gioti who will grow fungi in the lab, extract DNAs and RNAs (WP2) and will coordinate multiomic data integration in WP3 along with the HPC Zorbas team. A bioinformatics training event in 2023 is also planned, stay tuned!
-
Our paper (in collaboration with NTUA) elucidating a fungal symbiont’s ability to remove the persistent pollutant PCB29 in Science of the Total Environment https://www.sciencedirect.com/science/article/pii/S0048969721008858E. Nikolaivits, R. Siaperas, A. Agrafiotis, J. Ouazzani, A. Magoulas, Α. Gioti and E. Topakas. Functional and transcriptomic investigation of laccase activity in the presence of PCB29 identifies two novel enzymes and the multicopper oxidase repertoire of a marine-derived fungus. Science of the Total Environment, 775, 145818.
-
NEW IMBBC PROJECT FUNDED BY THIRA MUNICIPALITY
 THIRA: Shallow submarine volcanoes have complex geology and research into their hydrothermal systems is still at an early stage. These shallow submarine, high-energy, dynamic, hydrothermal systems are associated with significant and potentially hazardous volcanic activity. The Kolumbo volcano is the most active submarine volcano of the Mediterranean Sea. Thira is a collaborative project among the Municipality of Thira, the Hellenic Center for Marine Research (ELKETHE) and the National and Kapodistrian University of Athens (NKUA), which aims to monitor the underwater Kolumbo volcano using the R/V Philia and the Remote Operated Vehicle owned by HCMR.
THIRA: Shallow submarine volcanoes have complex geology and research into their hydrothermal systems is still at an early stage. These shallow submarine, high-energy, dynamic, hydrothermal systems are associated with significant and potentially hazardous volcanic activity. The Kolumbo volcano is the most active submarine volcano of the Mediterranean Sea. Thira is a collaborative project among the Municipality of Thira, the Hellenic Center for Marine Research (ELKETHE) and the National and Kapodistrian University of Athens (NKUA), which aims to monitor the underwater Kolumbo volcano using the R/V Philia and the Remote Operated Vehicle owned by HCMR. -
IMBBC participated to the IODP EXPEDITION 398 (December 2022 – February 2023). A number of samples were collected in order to characterize the living and fossilized subseafloor biological communities present and to document the sizes, genetic variabilities, and metabolic functions of subsurface ecosystems to 700 m depth.


















